What are the concentrations of free metabolites in cells?
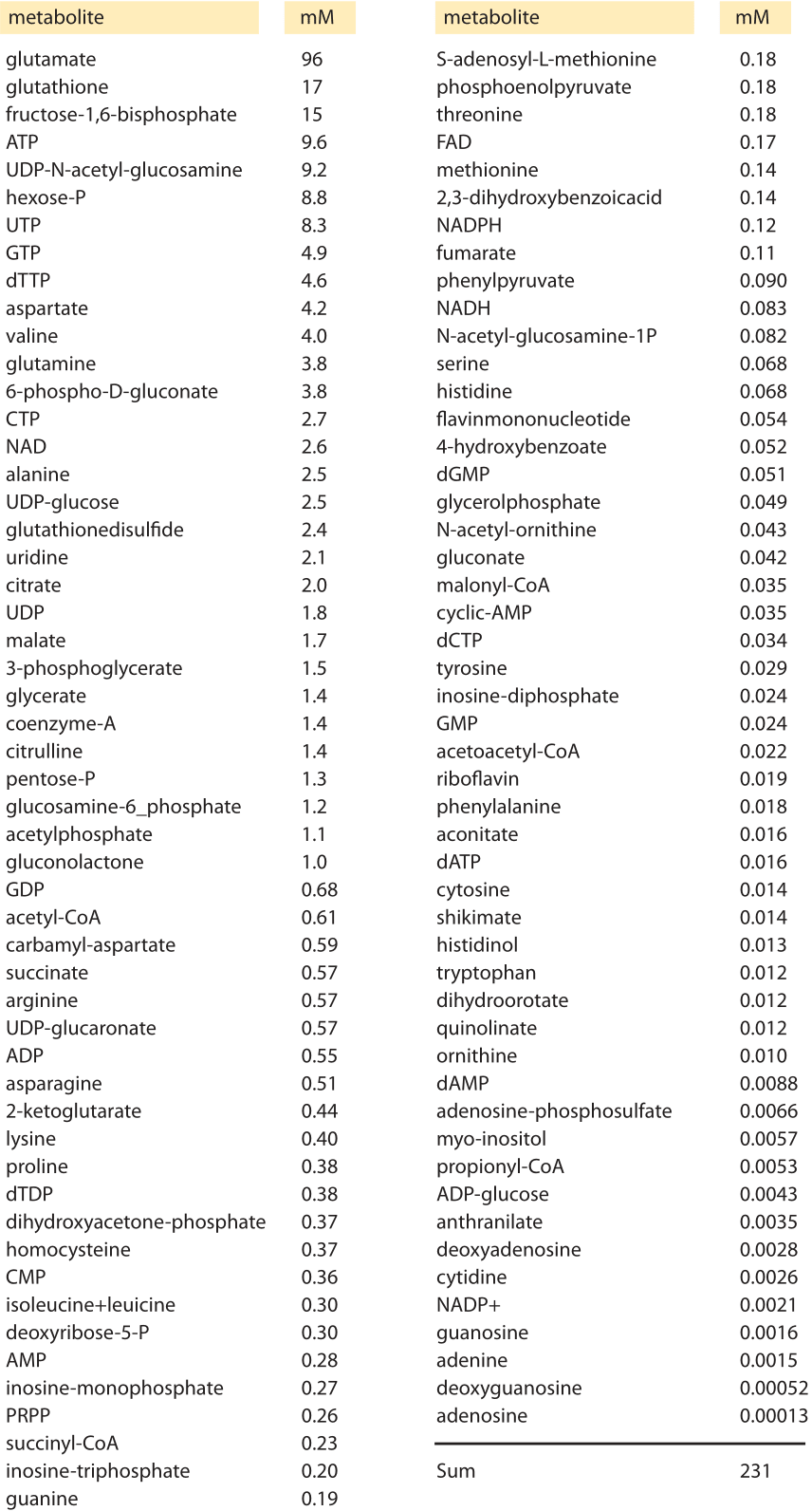
Table 1 Intracellular concentrations of the most abundant metabolites in glucose-fed, exponentially growing E. coli measured via mass spectroscopy. Adapted from: Bennett et al, Nature chemical biology, 2009
The cell’s canonical components of proteins, nucleic acids, lipids and sugars are complemented by a host of small metabolites that serve a number of key roles. These metabolites are broadly defined as members of the many families of molecules within cells having a molecular weight of less than 1000 Daltons. Recent measurements have made it possible to take a census of these metabolites in bacteria as shown in Table 1. Perhaps the most familiar role for these metabolites is as the building blocks for the polymerization reactions leading to the assembly of the key macromolecules of the cell. However, their biochemical reach is much larger than this restricted set of reactions. These metabolites also serve as energy sources, key activity regulators, signal transducers, electron donors and buffers of both pH and osmotic pressure.
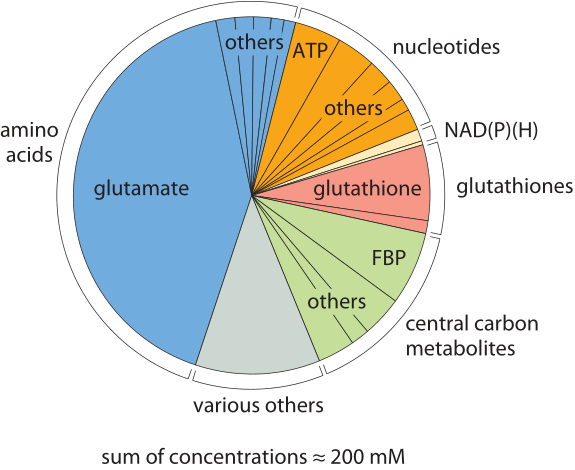
Figure 1: The composition of free metabolites for an E. coli cell growing on glucose. Metabolites are colored based on their functional group. In each category “other” refers to other metabolites in that category whose names are not shown due to small size. “misc.” refers to other metabolites not part of any of the other categories such as UDP-N-Ac-glucosamine. FBP stands for fructose-1,6-bisphosphate. (Adapted from B. D. Bennett, Nature Chem. Biol., 5:593, 2009.)
An inventory of which metabolites are present and at what concentrations is of great interest since it provides a picture of the stocks available to the cell as reserves for building its macromolecules. In addition, this inventory tells us which compounds are most ubiquitous and how we should think about the various chemical reactions (both specific and nonspecific) that they are part of. The concentrations of some metabolites are easy to measure whereas others are notoriously difficult. Thanks to advances in mass spectrometry the comprehensive survey of cell metabolite concentrations detailed in Table 1 and Figure 1 became possible. The table depicts the most abundant metabolites in E. coli during growth on M9 medium supplemented with glucose. These surveys do not include simple ions such as potassium and chloride which we discuss separately. Another key property beyond the concentration is the turnover time of the metabolite pool that we discuss separately in the vignette “What is the turnover time of metabolites and tRNAs?”.
The molecular census of metabolites in E. coli reveals some overwhelmingly dominant molecular players. The amino acid glutamate wins out in Table 1 at about 100mM, which is higher than all other amino acids combined as depicted in Figure 1. Our intuition and memory is much better with absolute numbers than with concentrations so we recall our rule of thumb that a concentration of 1 nM corresponds to roughly one copy of the molecule of interest per E. coli cell. Hence, 100 mM means that there are roughly 108 copies of glutamate in each bacterium. How many protein equivalents is this? If we think of a protein as being built up of 300 aa, then these 108 glutamates are equivalent to roughly 3 x 105 proteins, roughly 10% of the ≈3 x 106 proteins making up the entire protein census of the cell (BNID 100088). This small calculation also shows that the “standard conditions” of 1M concentration often employed in biochemical thermodynamic calculations are not realistic for the cell, as such a concentration will take up all the cell mass (or more for larger compounds). We note that glutamate is negatively charged, as are most of the other abundant metabolites in the cell. This stockpile of negative charges is balanced mostly by a corresponding positively-charged stockpile of free potassium ions (K+) which have a typical concentration of roughly 200 mM.
The second most abundant metabolite, glutathione, is the key regulator of the cell redox potential, strongly affecting protein structure by making and breaking sulfur bonds among cysteine residues. This key player is further discussed in the vignette “What is the redox potential of the cell?”. The third most abundant metabolite, fructose 1,6 bisphosphate is a central component of the carbon highway of the cell – glycolysis, with the fourth most abundant metabolite coming in as ATP, the main energy currency. Moving from the specific roles to the larger picture, one glaring feature revealed by the table is the broad range of concentrations found for these metabolites ranging from roughly 10-1 to 10-7 M. Given our rough rule of thumb that a concentration of 1 nM implies roughly 1 such molecule in the volume of an E. coli cell, this implies that the range of metabolite numbers inherent in these measurements is from as many as 108 copies of glutamate as noted above down to as few as 100 copies of the nucleoside adenosine, a million-fold range of concentrations.
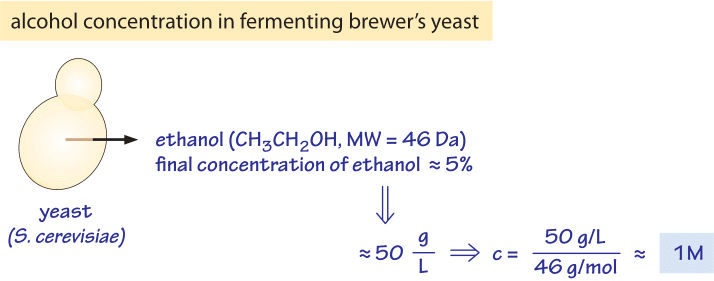
Figure 2: Concentration of alcohol produced by yeast. The ethanol content of wine, ≈5%, is equivalent to a concentration of about 1M, a very high osmotic pressure.
As seen in the table, the total concentration of free metabolites is on the order of 250mM. How can we put this number in perspective? One prominent example of where we have a feel for metabolite production is in the production of our favorite alcoholic beverages where ethanol is produced by yeast. Yeast produce beer and wine through fermentation of sugars to the alcohol ethanol. Fermentation results in 2-3 ATP molecules being produced per glucose molecule consumed, much less than the ≈30 that can be produced when using the TCA cycle. Yet, brewer’s yeast still prefers to perform fermentation even when oxygen is available for the TCA cycle. One explanation for this odd behavior from the perspective of energy utilization is that fermentation with its associated excreted byproducts creates an environment that is not suitable for other organisms that inhabit the same niche. As such, one might speculate that by producing this alcohol, yeast effectively forbids bacteria to grow nearby. In Figure 2 we provide a schematic of the numbers associated with fermentation which serve as the basis for this speculative mechanism that awaits rigorous experimental examination. The alcohol content in beer is typically ≈4% and in wine ≈12%. Ethanol (H3CCH2OH) therefore has a concentration of 40 g/l and 120 g/l, respectively. With a molecular weight of 46, we show in the figure that 5% is equivalent to ≈1 M concentration. Many bacteria are not able to grow in the presence of such high concentrations of alcohol (K. Tamura et al., FEMS microbiology letters, 99:321, 1992) which affects what is termed the “fluidity” of the cell membrane as well as the contents of the cell interior (ethanol is a small molecule to which the membrane is partially permeable). The brewers’ yeast is adapted to these high ethanol concentrations and also to the low pH which both inhibit the growth of most bacteria. Indeed after the completion of the fermentative phase, yeast move to a phase of respiration making use of the extra energy capacity of ethanol with relatively little competition. It has been suggested that this is the reason that yeast choose this growth strategy (J. Piskur et al, Trends in Genetics, 22:183, 2006). We note though that this speculation on yeast growth strategy has been criticized (D. Molenaar et al, Mol. Sys. Biol., 5:323, 2009) as not being an evolutionary stable strategy against “cheater” mutants that will not produce ethanol but would still enjoy the lower competition and thus could penetrate and overtake the ethanol producing population.
In summary we return to the table on the abundance of metabolites. Why are some metabolites present in the cell at such high abundance while others are present at such minute concentrations? The starting point of an analysis of this question should focus on the costs and benefits of maintaining each of these metabolites at a given concentration. It is of interest to learn whether these concentrations are dictated by some adaptation that has been sculpted by evolution. Rationalizing the top metabolites such as glutamate, FBP and glutathione seems like a natural place to begin addressing this challenge. Surprisingly, we have been unable to find rigorously articulated and experimentally corroborated explanations for the relative concentrations of different metabolites, a befitting future challenge for cell and systems biologists.

